cancelling an ENA record which has been released to the public) will not be possible. Given the split of meta-data and data files between ArrayExpress and ENA, once your submission is fully processed, it is a lengthy process to modify/update it. Depending on the file format, it will either be stored at ArrayExpress or the ENA. BAM alignment files, differential expression data, expression values linked to genome coordinates, etc). You can also send us processed data (i.e.
ArrayExpress will transfer the raw data files to the ENA for you so you do not need to submit those files separately to the ENA.
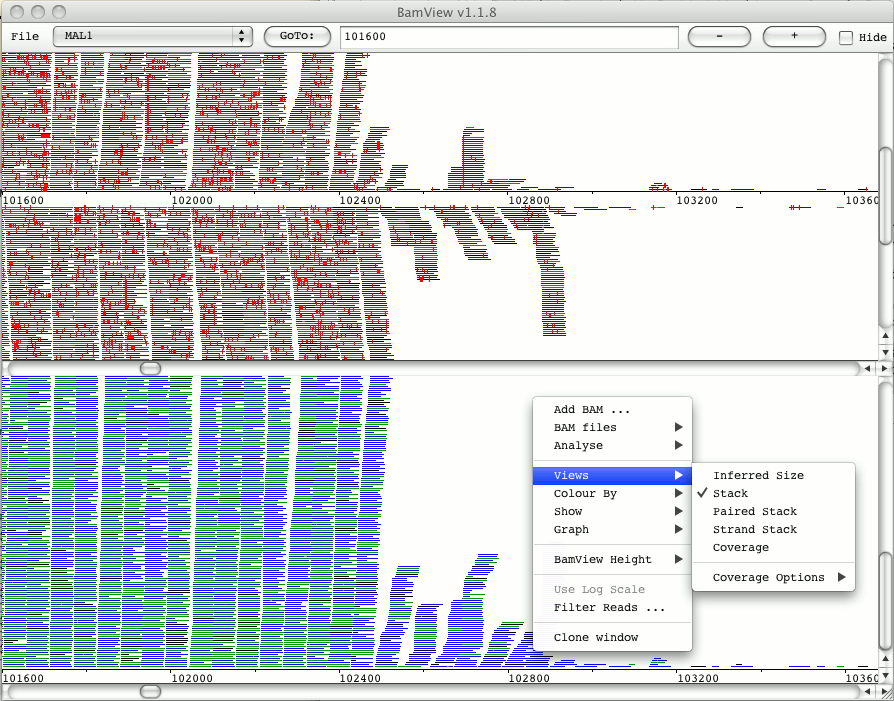
#.bam file format sequencing data archive
fastq files) are stored at the Sequence Read Archive (SRA) of the European Nucleotide Archive (ENA). The meta-data about your experiment will be stored at ArrayExpress, and the raw data files (e.g. Submissions without raw data files will not be accepted unless there are exceptional circumstances. experiment description, samples and their attributes, all protocols used) and the raw data files see the submission guide below. To submit to ArrayExpress, all you need to do is send us meta-data for your experiment (e.g.
If you have potentially identifiable human data, please see below. Once we have finished processing your raw reads, submit the assembled transcriptome file (often in fasta format) directly to the European Nucleotide Archive.

ArrayExpress accepts submissions of functional genomics data generated using high throughput sequencing (HTS) assays like RNA-seq and ChIP-seq, mostly from non-human and human non-identifiable samples, with the following exceptions:
